Rewinding Evolution: How Ancient Antibiotics Could Help Fight Modern Superbugs
- Jake Klardie
- Apr 17
- 1 min read
Antibiotic resistance is one of the biggest challenges we face in modern medicine. Bacteria are evolving faster than we can develop new drugs making once treatable infections increasingly dangerous and even deadly. But what if the solution to this problem isn’t entirely new and rather, very old.
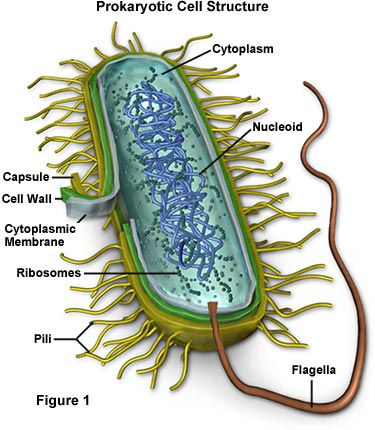
A recent study published in Nature Communications explored that idea by resurrecting ancient antibiotic molecules. In this research, scientists investigated a class of antibiotics called glycopeptides which target lipid II, a molecule essential for bacterial cell wall synthesis. Lipid II is a particularly attractive target because by disrupting it, it prevents bacteria from building a protective outer structure ultimately leading to cell death.

The researchers used computational biology and synthetic biology to reconstruct an ancestral antibiotic molecule called paleomycin. By analyzing genetic sequences and enzyme pathways, they were able to predict how ancient bacteria produced the compound. They then synthesized it in the lab and confirmed that it still had antibacterial activity.
This discovery is exciting because ancient antibiotics may work differently than newly synthesized ones. Because bacteria haven’t encountered these ancient bacteria in millions of years, they may not have developed any form of resistance to them unlike their modern counterparts. This provides scientists with a new strategy, instead of searching for novel bacteria with new antibacterial mechanisms, they can mine evolutionary history for forgotten antibiotics.

This research opens a new direction in antibiotic discovery. While traditional approaches often struggle due to the speed of bacterial adaptation, targeting conserved molecules like lipid II and doing so with structurally unique compounds may slow the development of resistance.

Comments